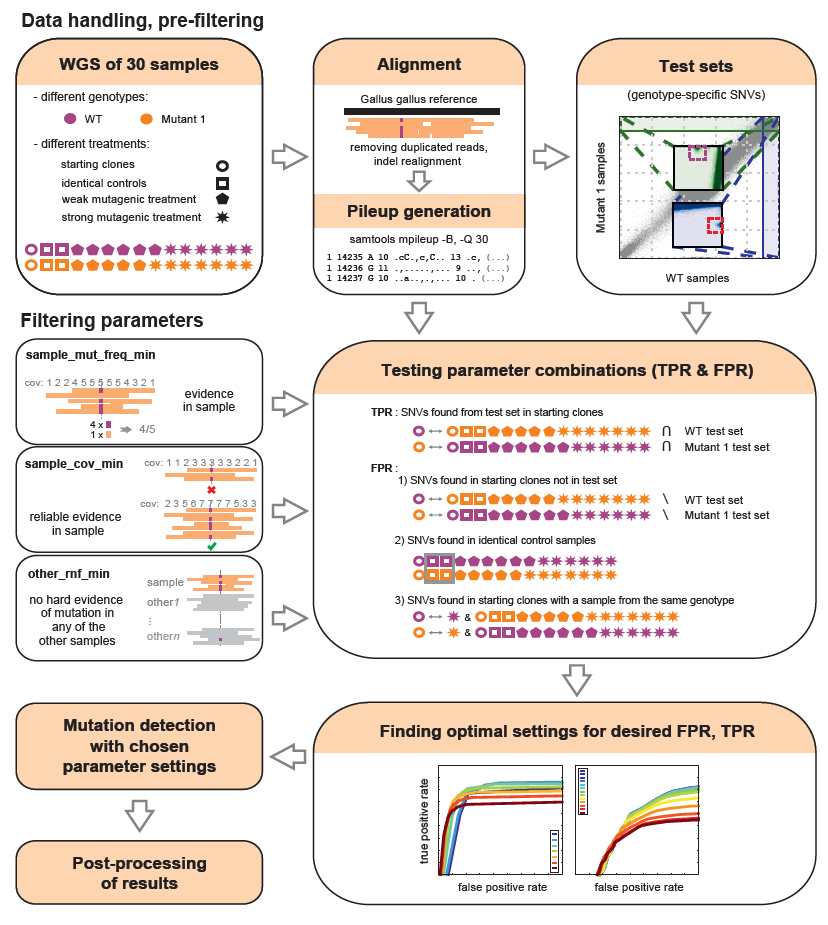
IsoMut is a tool designed for experimental scenarios whenever unique mutations (single nucleotide variations or insertions/deletions) are sought in multiple isogenic samples. The software uses information gathered from all the available samples to rule out false positive mutations with great efficiency. On a 30 sample dataset at a mean sequence coverage of 20, IsoMut could detect over 90% of true heterozygous single nucleotide variations with no more than 25 false positives per gigabase of genome sequence. A general overview of the method can be seen on the figure below. For more details and guidelines on usage, see Publications. To download IsoMut, see the Download page.